Evidence-Based Design and Evaluation of a Whole Genome Sequencing Clinical Report for the Reference Microbiology Laboratory
Abstract | Paper | Talk | Poster | Additional Materials |
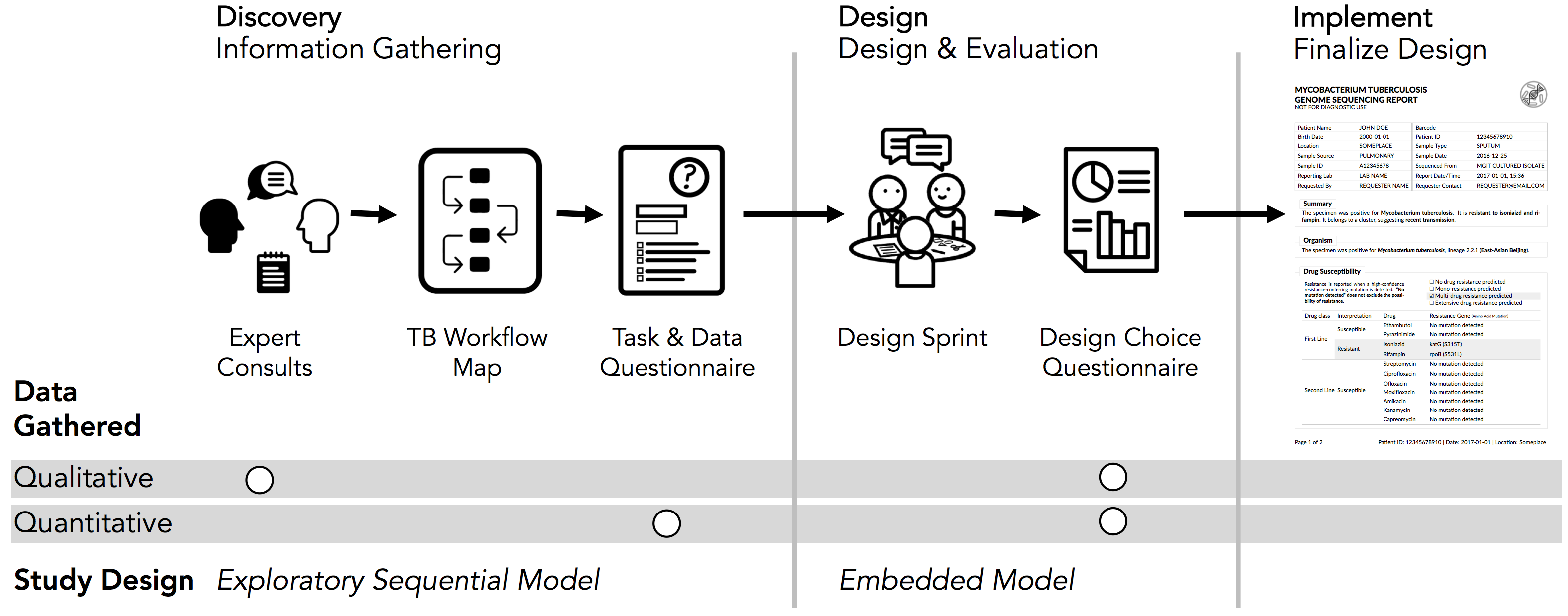 Fig. 1. An overview of our appraoch, which interleaved mixed-methods designs with infovis Design Study Methdology.
Fig. 1. An overview of our appraoch, which interleaved mixed-methods designs with infovis Design Study Methdology. Abstract
The outputs of this project primarily target domain specialistis in microbial genomics. However, we have relevant findings for Vis researchers too. Thus, two abstracts are provided below to address our message to the infovis and microbial genomics communities.Vis Specialist Abstract
Design study methodologies developed by infovis researchers are not adopted beyond this community despite a growing repertoire of visualization tools created by domain specialists. While better knowledge translation between infovis and domain specialists is needed, it is also arguable that visualization tools, which can have many complex interacting parts, can make it challenging to demonstrate the design study methodology in action. Here, we present our application of a design study methodology to a simpler problem of information design in a tuberculosis (TB) clinical report that presents results derived from TB whole genome sequencing (WGS). Clinical reports are a common and foundational element of medical and public health practice and as such are a good, and relatively simple, application context in which to demonstrate the value of applying a design study methodology. Using an existing TB clinical WGS report as a base, we collected relevant tasks and data, linked those to alternative report designs, and finally compared those alternatives to the original report with stakeholders. The evidence gathered through our project demonstrated how the original, ad hoc, report design contained elements that were unnecessary, difficult to interpret, or insufficient. We also demonstrated how a number of procedural constraints around current reporting practices, such as how stakeholders received reports and how much time they had to review them, affected the report’s design. Taking into consideration the evidence gathered and regulatory guidelines, we produced a new TB WGS clinical report that is currently under more detailed assessment and awaiting deployment. By using a simple and relatable design challenge, providing a concrete framework for data gathering and analysis, and through participatory engagement of the relevant domain specialist community, our work aimed to introduce infovis evidence-based design methodologies to the health sciences community.Domain Specialist Abstract
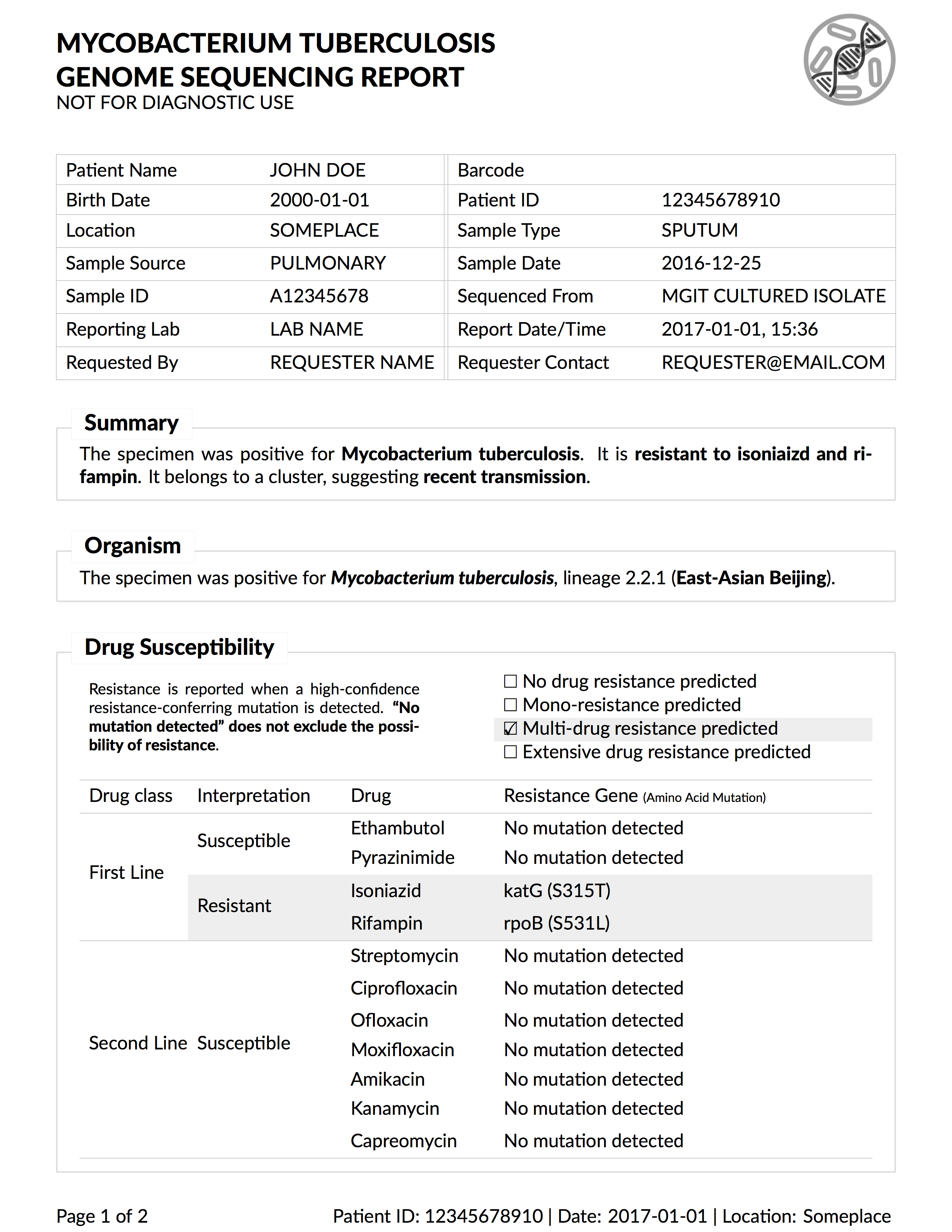
Background: Microbial genome sequencing is now being routinely used in many clinical and public health laboratories. Understanding how to report complex genomic test results to stakeholders who may have varying familiarity with genomics – including clinicians, laboratorians, epidemiologists, and researchers – is critical to the successful and sustainable implementation of this new technology; however, there are no evidence-based guidelines for designing such a report in the pathogen genomics domain. Here, we describe an iterative, human-centered approach to creating a report template for communicating tuberculosis (TB) genomic test results.
Methods: We used Design Study Methodology – a human centered multi-stage approach drawn from the information visualization domain – to redesign an existing clinical report. We used expert consults and an online questionnaire to discover various stakeholders’ needs around the types of data and tasks related to TB that they encounter in their daily workflow. We also evaluated their perceptions of and familiarity with genomic data, as well as its utility at various clinical decision points. These data shaped the design of multiple prototype reports that were compared against the existing report through a second online survey, with the resulting qualitative and quantitative data informing the final, redesigned, report.
Results: We recruited 78 participants, 65 of whom were clinicians, nurses, laboratorians, researchers, and epidemiologists involved in TB diagnosis, treatment, and/or surveillance. Our first survey indicated that participants were largely enthusiastic about genomic data, with the majority agreeing on its utility for certain TB diagnosis and treatment tasks and many reporting some confidence in their ability to interpret this type of data (between 58.8% and 94.1%, depending on the specific data type). When we compared our four prototype reports against the existing design, we found that the majority (86.7%) of design comparisons, participants preferred the alternative prototype designs over the existing version, and that both clinicians and non-clinicians expressed similar design preferences. Participants articulated clearer design preferences when asked to compare individual design elements versus entire reports. Both the quantitative and qualitative data informed the design of a revised report, which is available online as a LaTeX template.
Conclusions: We show how a human-centred design approach integrating quantitative and qualitative feedback can be used to design an alternative report for representing complex microbial genomic data. We suggest experimental and design guidelines to inform future design studies in the bioinformatics and microbial genomics domains, and suggest that this type of mixed-methods study is important to facilitate the successful translation of pathogen genomics in the clinic, not only for clinical reports but also more complex bioinformatic data visualization software.
Paper

Talk
Anamaria Crisan will present on this project in the From Pipelines to Pixels: NGS Data Integration (Reporting, QC/QA, Accreditation, Training) and Visualization session at 4:00 PM EST on October 9th, 2017 at ASM NGS Pipelines Conference.[Talk]
Additional Materials
Report Design Template
A LaTeX template of the report design is avialable on Overleaf or through a direct download.Project Data and Analysis
Qunatitative data from the different questionnaires has been made available on github
Last modified: Oct 8, 2017.