Michael DiBernardo
Poutanen SM et al. New England Journal of Medicine 2003 May 15;348(20):1995-2005.
http://content.nejm.org/cgi/content/full/348/20/1995
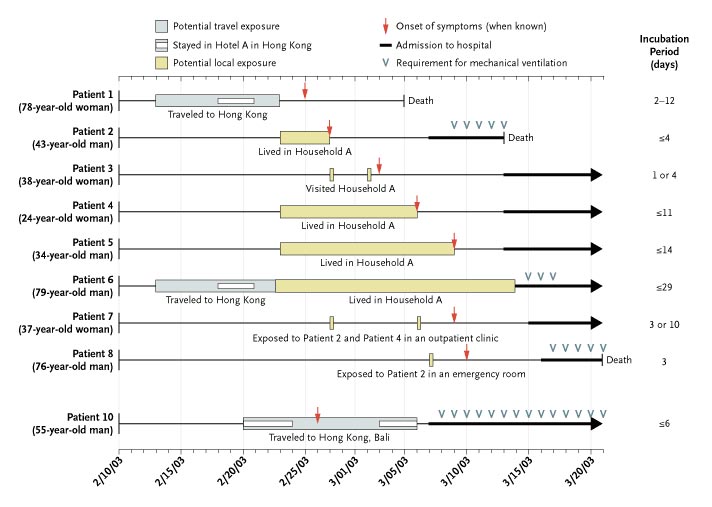
This article describes the relationships between the Canadians that contracted SARS in 2003, and infers properties of the disease (mechanism of transmission, time of incubation, lethality, etc.) from this data. The provided diagram is an attempt to summarize the important characteristics of these relationships over time.
I believe that this diagram has a single property that makes it a useful visualization of the data: The organization of the picture makes it very easy to pick out possible answers to many initial questions one may have about the disease because the colors and symbols chosen are distinctive enough to allow you to discard all of the details that are nonessential to your question. For instance, if you wish to know the extent of exposure that was necessary before symptoms were displayed, one can pay attention only to the boxes and the vertical arrows. If you would like to have a rough idea of the effects of early diagnosis on the rate of survival, one can compare the length of the line between the vertical arrow and the bold horizontal arrow between those patients who died and those who lived. Unfortunately, this ease of abstraction may also encourage you to make inferences that are confounded with patterns that you are purposely ignoring.
Pietrokovski S. Searching databases of conserved sequence regions by aligning protein multiple-alignments. Nucleic Acids Research 1996 Oct 1;24(19):3836-45
http://bioinformatics.oxfordjournals.org/cgi/content/full/21/18/3596
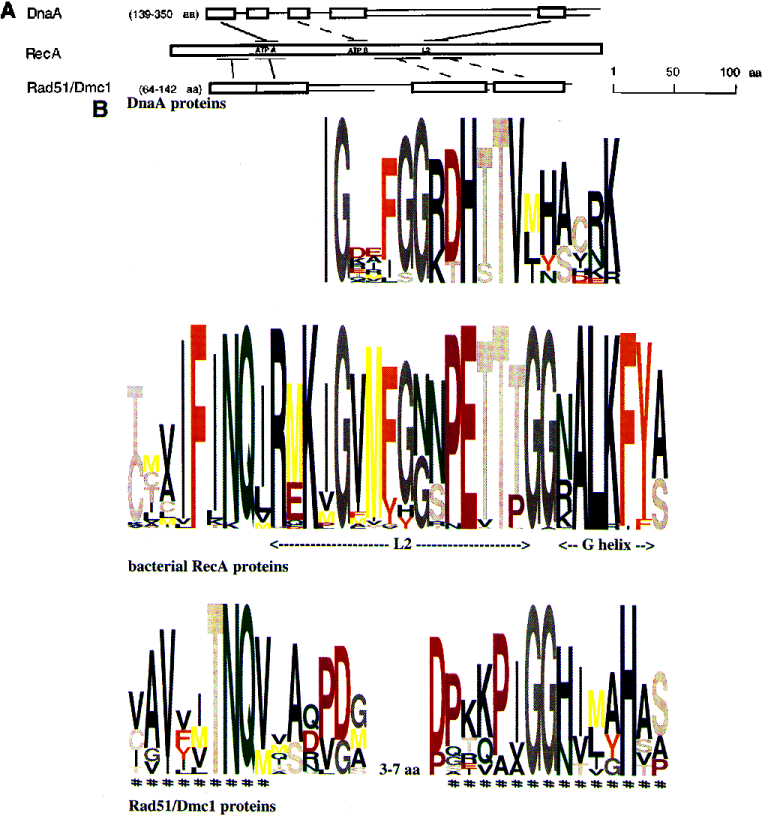
While I'm picking on one publication in particular here, logo diagrams are often used in bioinformatics to describe the positional frequency of amino acids across short stretches of functionally relevant sequence called motifs or domains.
I find these diagrams to be horrible in many ways. They are aesthetically very unpleasant, since twenty different colours are used to distinguish the twenty different single-letter amino acid codes; this is done because the less frequent residues may have codes that look similar (P and R, for example). This distraction also makes it difficult to distinguish any useful information about a particular position except for the most frequent residue in that position, which could have been elucidated much more clearly in a simple table or bar chart of probabilities in any case. In cases when there is no single dominant residue, it is difficult to gauge the relative frequency of one residue with respect to another. Finally, in many of these diagrams (including this one), extremely infrequent residues are not even included in the visualization, and so each column terminates at a different height. This is nonintuitive: One would expect the sum of all of the frequencies in each column to add to 1.0, and thus for all of the columns to be of equal height.