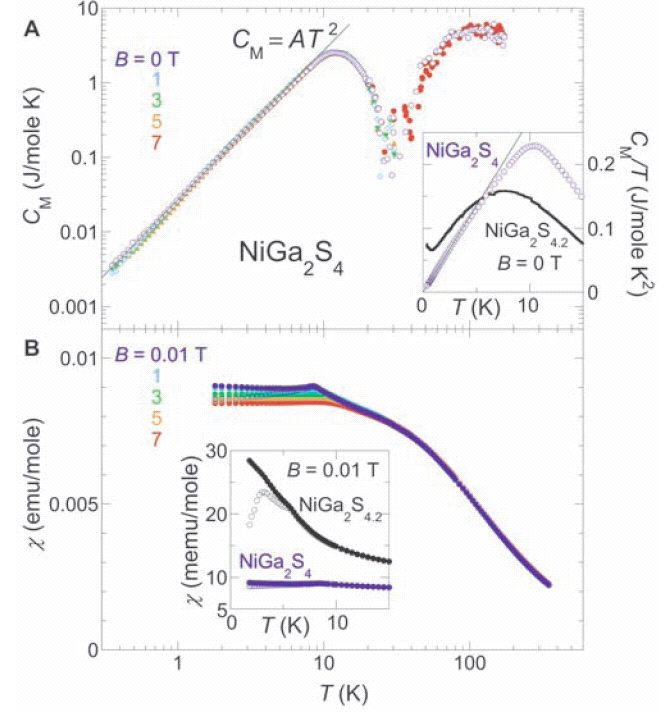
The A section of this diagram shows the specific heat of NiGa2S4 versus temperature under the influence of a magnetic field. The thin solid black line represents the theoretically predicted specific heat. The coloured circles represent the observed specific heat. Each magnetic field strength B is represented by a different colour. The B section and the inset diagrams will not be the focus of this discussion. The important information in the diagram is the effect that each field strength (B = x) had on NiGa2S4, compared to the absence of a magnetic field (B=0). Unfortunately, bad visualization decisions hide rather than accentuate this comparison.
Colour choices cause a number of problems in this visualization. The red data points leap out at the viewer, but B=7 is an uninteresting case that deserves no more highlighting than the other field strengths. The light orange colour of B=5 vanishes into the background. It is hard for the viewer to pick out more than 3 or 4 orange data points with a casual glance. Another problem is the layering of the data. It is unclear why B=0 and 1 have hollow circles while B=3,5, and 7 have solid circles. The hollow circles sink to the bottom of the visualization. This is particularly bad for B=0, as our important baseline data set is hidden at the bottom of the pile. The choice of circles to represent the data points instead of smaller dots is also dubious. The eye is immediately drawn to regions where the data points look like individual circles instead of a continuous line. These regions are just artifacts of the logarithmic scale and data acquisition timing and not actual areas of interesting data. Finally it is clear that too much data is being forced into a small space. The inset diagrams are confusing as they contain different scales, data ranges, axis values, and refer to different experiments.
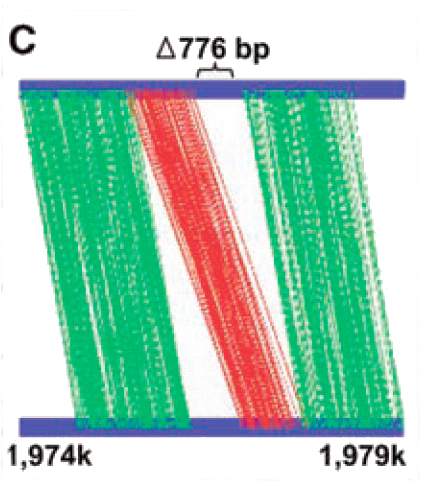

The second visualization is from a paper describing a new low cost DNA sequencing technology. The visualization shows the error that occurred when the low cost sequencing of E. Coli was compared to standard sequencing methods. The top blue line shows the genomic positions in the low cost sequence. The bottom blue line shows the genomic positions of the standard sequence. Correct sequencing is shown in regions connected with green lines. Red lines show aberrant sequencing. The visualization highlights the absence of a 776 base pair sequence in the low cost method.
The colour choices are strong. Western culture is very familiar with the stoplight analogy with green indicating good proceed and red indicating stop error. The connecting lines make it clear that it is the relative sequencing genes that is important and not the absolute position. Compared to the thickness of red and green lines, the blank white space makes gaps in the sequencing data easy to spot. Finally the important gap of 776 base pairs is explicitly highlighted. This visualization's one major failing is that all labels and key information is described in a paragraph below the visualization rather than on the visualization itself.