- a cross between biology & info sciences
- IT/CS tools for biological problems
(why not biological tools for IT/CS problems?)
- CS research motivated by biological questions
- information processing in biological systems
- &c.
Webpage: http://www.cs.ubc.ca/labs/beta/courses/bioinf536a.html
There will be a box in the Reading Room for articles, &c.
The project will probably be done in groups of about 3.
Course outline: 7 modules:
I. Intro / Basics
II. Sequence Alignment
III. Phylogenetic Analysis
IV. RNA / Protein Structure
V. Gene Finding / Sequence Annotation
VI. Gene Expression / Regulatory Networks
VII. Biomolecular Computing
The basic genetic process:
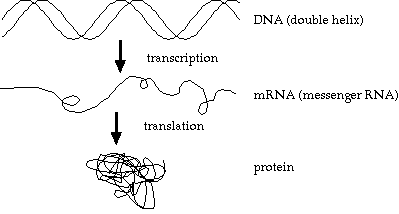
Unfortunately, this is an overly simplistic view. Details of the processes are usually omitted in bioinformatics texts. If aimed at biologists, it’s assumed they don’t need the details. If aimed at computer scientists, it’s assumed they don’t want the details.
So, here is an improved version (still not perfect):
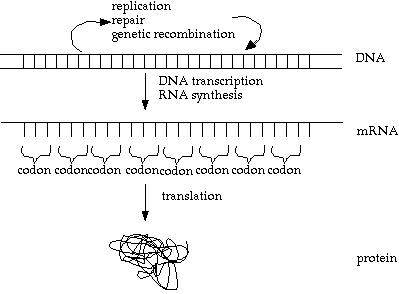
Note however, that something called 'reverse transcriptase' reverses the info flow, allowing information on a strand of mRNA to be inserted into the DNA. This is how viruses such as HIV can invade DNA.
DNA Structure:
So what does each strand of DNA look like?
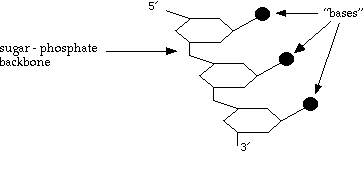
The bases can be: T thymine
A adenine
C cytosine
G guanine
Note that the ends are chemically distinct, and are labelled
 and
and 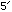 by convention (this imposes an implicit order on the chain)
by convention (this imposes an implicit order on the chain)Also important is the fact that C & G form hydrogen bonds with each other to link between the two strands, as do A & T. Other pairings are unlikely.
By convention, DNA is written from the 5’ end to the 3’ end, e.g.:
5' - CGATT . . . . T-3'
The double helix formed by the two strands is actually asymmetric, and has a “major groove” and a “minor groove”, which becomes important for some interactions.

Protein Structure:
Proteins also have a backbone, formed of polypeptides, with a wide variety of side groups (there are 20 amino acids normally used, each of which has a 3-letter and a 1-letter abbreviation). These side groups have various chemical properties, which are important to the structure and function of the protein.
As with DNA, the ends are chemically different (in this case, an N-terminus and a C-terminus).
For most proteins, the function is dictated by the three-dimensional structure, which in turn depends on the sequence of amino acids along the backbone (massive simplification here . . .).
By convention, proteins are also written left to right, e.g.:
N - DKNSE . . . . D - C
or N - ArgLeuLysArgArg . . . .Pro - C
One common motif in proteins is the "leucine zipper", in which repeated leucines in two places line up to form a rigid finger that fits neatly into the major groove of DNA.
Proteins fall into a number of general categories:
enzymes (catalysts for chemical reactions)
regulatory (control the action / generation / destruction of others)
transport
storage
contractile / motile
structural
protective
exotic
RNA Structure:
RNA is the (usually) single-stranded analogue of DNA, with a slightly different backbone, and with a base called uracil (U) substituted for thymine (T). The backbone is less stable than that of DNA. Note that cytosine is slightly unstable, and can actually degenerate into uracil spontaneously.
As to structure, RNA generally floats around as a single strand, but can coil up and interact with itself in odd ways - this is significant.
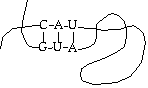
One of the results of this is that RNA can form functional shapes, in much the same way as protein does. It is currently believed (the "RNA world hypothesis") that RNA is the original self-replicating molecule, and that both proteins and DNA evolved later for greater efficiency. Thus, although the simple model treats RNA as an intermediary, it may have other functions. For example, there is a known RNA molecule whose function is to chop up other strands of RNA.
DNA Transcription:

Suppose we need to unzip the helix, and make a copy of a specified strand. What actually happens is that a hole is made (by breaking the bonds between base pairs, and we walk along the chain, matching up little bits of RNA as we go:
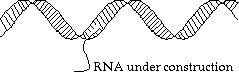
Protein Assembly:
To translate a strand of mRNA to a protein, an object called a `ribosome' attaches to a strand of mRNA, and translates it into a protein, 3 bases at a time. The ribosome itself consists of RNA and protein components that self-assemble into a functional complex that perform transcription (i.e. the ribosome).
Actually, this description of transcription is a simplification: multiple ribosomes can attach to the same mRNA strand, pipelining the protein construction process. And in some organisms (some bacteria, for example), ribosomes can attach to a strand of mRNA while it is being transcribed from the DNA.
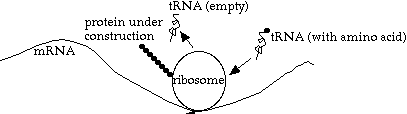
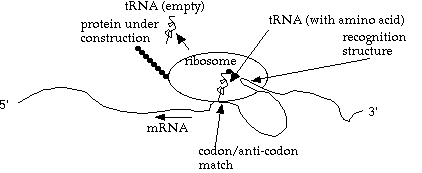
What happens inside the ribosome? A strand of tRNA (transfer RNA), with one amino acid attached, moves in. If it has the complementary codon (3 base group) to the "current" codon on the mRNA, then the amino acid is linked onto the protein chain, the ribosome advances one codon, and the now-empty tRNA is released to go find another (identical) amino acid.
Note that each tRNA molecule is customized for one particular type of amino acid, and one anti-codon (the opposite of the codon it matches to on the mRNA).
Codons:
With 4 possible amino acids, and three bases to a codon, there are 64 possible codons, so the coding is degenerate(i.e. there are usually several codons for a given amino acid).
For example: UUU = Phe
CCU = Leu
In addition, there are three "stop" codons, used to mark the end of the protein. These stop codons are called "amber", "ochre" and "opal".
The coding for amino acids is largely consistent across organisms, which argues for a single point of origin for this mechanism. Also, the third base in the codon is the least significant - for example UUA, UUC, UUG, and UUU all refer to Phe. This corresponds to the observation that, in the ribosome, the third hydrogen pair bond between the codon and anti-codon is the weakest.
Also remember that this process is not 100% accurate.
DNA Splicing:
In more complex organisms, the individual 'genes' that encode proteins are often broken up in the DNA, so there is a stage of transcription called 'splicing', in which the relevant chunks are chopped up and reassembled to get the 'real' gene. This means that looking for a particular protein's gene in DNA is tricky (i.e. it is not a simple string search).
Types of RNA:
We have already seen mRNA and tRNA. There is also rRNA - ribosomal RNA, because the ribosomes are principally constructed from RNA (looks like bootstrapping to me . . .).
In E. coli, for example, the RNA in a cell breaks down as follows:
80% rRNA
15% tRNA
5% mRNA
Of these forms, mRNA is actively broken down in the cell, in order to facilitate gene product regulation (if it stayed around forever, it would be hard to stop making a particular gene product). tRNA, on the other hand, is recycled: when the amino acid is detached from tRNA during transcription, the tRNA floats off to find another molecule of the amino acid.
A Weird Example:
An exception to the rule that codons are consistent across species is "procaryotic selenocysteine insertion", in which an extra amino acid (selenocysteine) can be inserted into a protein when the UGA (amber?) stop codon is met
What happens here is that if a suitable chunk of the mRNA strand (further downstream: i.e. towards the 3' end) matches a structure built into the ribosome, the effect of the codon/anti-codon match is modified to substitute a selenocysteine instead of stopping.
If a subsequent amber stop codon is encountered, it will revert to the stop meaning. (It was not clear whether this insertion can happen multiple times in the same protein). Also note that the recognition structure may actually be composed of codons that have meaning for the protein!
In any event, this changes the "grammar" of RNA-protein translation from context-free to context-sensitive!
A similar situation occurs when a 'pseudoknot' structure is used to force a UAG codon (another stop codon) to act as a GAG codon.